Introduction
Materials and Methods
Sampling
Scanning electron microscopy (SEM)
PCR and MinION sequencing
Sequence analysis and bioinformatics
Results
Environmental parameters
Diatoms on microplastics
Diatoms are relatively more abundant on microplastics compared to the surrounding seawater
The microplastic contain a distinct composition of pennate diatoms
Discussion
Introduction
Marine plastic pollution is a growing environmental threat as the volume of plastic waste entering oceans continues to rise (Borrelle et al. 2020). Over time, larger debris fragments into microplastics (≤5 mm), which have been detected in marine ecosystems worldwide. The semi-enclosed Mediterranean Sea, surrounded by densely populated coastlines, is a known hotspot for microplastic accumulation (Cózar, 2015, Terzi et al. 2024). Surface waters off the Israeli coast have been reported to contain extremely high microplastic concentrations – up to 324 particles per cubic meter in some locations (van der Hal et al. 2017). Recent analyses revealed that the floating microplastic in the Mediterranean Sea is composed mainly of Polyethylene (PE), Polypropylene (PP) and polystyrene (PS) with high percentages of transparent particles (van der Hal et al. 2017, Marsay et al. 2023).
Plastic debris, including microplastic particles serve as stable surfaces, supporting the development of complex communities, including bacteria, protists, fungi, algae, and sessile invertebrates (Amaral-Zettler et al. 2020). The plastic community is taxonomically distinct from the planktonic community in surrounding seawater (Davidov et al. 2020, Davidov et al. 2024). It is therefore regarded as a separate ecosystem termed “plastisphere” (Zettler et al. 2013). While the type of plastic polymer influences the initial composition of the microbial colonizers (Marsay et al. 2022) this effect diminishes over time as the particles become uniformly coated with a mature biofilm layer (Davidov et al. 2025). Furthermore, it was established that the plastisphere communities strongly influenced by geographic location and seasonal variation. Spatiotemporal conditions affect key environmental parameters, including temperature, salinity, nutrient concentrations, pH, light availability, dissolved oxygen and CO₂ levels, as well as water turbulence. Fluctuations in these seasonally variable factors play a major role in shaping the taxonomic composition of the plastisphere microbiome (Oberbeckmann et al. 2014, Oberbeckmann et al. 2016, Marsay et al. 2023, Davidov et al. 2024).
Marine diatoms are a diverse group of unicellular, photosynthetic algae characterized by their intricately patterned silica cell walls (frustules). Diatoms play a central role in the oceanic carbon and silicon cycles. They are key primary producers in ocean ecosystems, contributing an estimated 40–50% of the total marine primary productivity (Rousseaux and Gregg 2013, Sarthou et al. 2005) and an estimated 20–25% of the global primary production, carbon fixation, and atmosphere oxygen release (Field et al. 1998, Serôdio and Lavaud 2022). Diatoms are found throughout the photic zone, from coastal waters to the open ocean, and can exist as free-floating (planktonic) or surface-attached (benthic) forms. Their ability to rapidly respond to nutrient availability makes them important indicators of environmental change.
In spite of their ecological importance and high abundance on plastic surfaces, their role within the plastisphere remains poorly understood. In particular, studies investigating diatom colonization, diversity, and ecological interactions on plastic debris in the Mediterranean Sea are still limited. Following our preliminary results and to address the above gaps, we re-analyzed our 18S rRNA metabarcoding dataset and used scanning electron microscopy (SEM) to further investigate the composition of diatom assemblages within the EMS plastisphere.
Materials and Methods
Sampling
The sampling was performed 1–1.2 km off the coast of Israel in the EMS (32°06'10.1"N, 34°46'51.2"E). Samples were collected seasonally during 2020–2021 from the water surface. Collection was done using a manta net with a mesh hole size of ~300 µm which was towed for 20–30 minutes at 2–3 knots along a fixed transect. Environmental parameters were taken along with the samples (Table 1). The net was deployed from a research vessel (Mediterranean Explorer, https://www.ecoocean.org/en/) and towed parallel to the coastline at 2–3 knots for 20–30 minutes, with three repetitions along the same transect. Particles recovered from the collection cup (at the tip of the net) were rinsed with autoclaved filtered (0.22 µm) artificial seawater (40 ppt, Red Sea salt, Red Sea Inc.). For DNA metabarcoding analysis of the plastic microbiome, a minimum of 10 microplastic particles were randomly selected from each sample. To analyze the water microbiome, 0.5 L of whole seawater was filtered through a 0.22 µm polyethersulfone membrane (Millipore) using a 20 L/min pump (MRC). Environmental parameters of the water column were measured using a CTD device at a depth of 0.5–1.5 meters.
Scanning electron microscopy (SEM)
Microplastic particles were fixed for 2–5 hours at room temperature in a solution of 1% glutaraldehyde and 4% paraformaldehyde (PFA), then washed three times with distilled water (5 minutes each) and stored in 50% ethanol-PBS at −20°C. Prior to imaging, samples were dehydrated in a graded ethanol series (50%, 70%, 85%, 95%, and 100%), with each step lasting 10 minutes, followed by three 15-minute incubations in 100% ethanol. Samples were air-dried for at least 5 hours under a fume hood, sputter-coated with a 10 nm platinum/gold layer (Quorum Q150T ES), and imaged using an Ultra-High-Resolution FE-SEM (Maia 3, Tescan) at 3–7 kV.
PCR and MinION sequencing
The amplification and sequencing procedures were the same as outlined in (Davidov et al. 2024). DNA was extracted from the microplastic and water sample using the DNeasy PowerWater Kit (Qiagen). 25–75 ng of template DNA were used for the downstream amplification reactions (PCR). The PCR products were cleaned with a QIAquick-PCR Purification kit (QIAGEN) to meet the criteria of the MinION nanopore library preparation protocol. For the barcoding of eukaryotes including most diatoms, the V4–V5 region of the 18S rRNA gene (18S in short) was amplified with 566F (5’ CAGCAGCCGCGGTAATTCC 3’) and 1289R (5’ ACTAAGAACGGCCATGCACC 3’) primers to generate ~0.7 kb long sequences according to (Davidov et al. 2020). The 18S sequencing libraries were prepared using the Native barcoding (EXP-NBD104) and ligation (SQK-LSK109) kits and protocols. The 18S multiplexed libraries were loaded onto separate MinION Nanopore Spot-on flow cells (FLO-MIN106D, version R9) and sequenced for 72 hours or until reaching ~7 Giga nucleotides (~4 M reads). Base-calling for all libraries was carried out by the Guppy base calling software 3.3.3, using the MinKnow program with the “high accuracy” option. Raw default “pass” quality reads were subjected to further analysis.
Sequence analysis and bioinformatics
The bioinformatics procedures and sequence analyses steps were performed according to (Davidov et al. 2024) using MetONTIIME pipeline and QIIME2 plugins (Bolyen et al. 2019). Reads were demultiplexed and trimmed from adaptors and PCR primers. Subsequently, sequences underwent quality filtering (quality ≥ 10) and length filtering. The length filtering was determined by read length histograms, with amplicon lengths falling within specific ranges of 650–1000 nucleotides. Clustering and taxonomic classification were performed separately for each barcode. Sequences were clustered into consensus sequences with default MetONTIIME pipeline parameters. Taxonomy was assigned to the 18S amplicons using BLAST against the Silva132 database (Quast et al. 2012), with a 90% identity threshold. All downstream analyses were based on relative abundance values, calculated by dividing the number of reads for each taxa in a sample by the total reads count. Only samples containing more than 10,000 reads were chosen for relative abundance analyses. The selected samples (>10 k reads) were all rarified to 10 k reads using MicrobiomeAnalyst 2.0 web platform (Lu et al. 2023).
Results
Environmental parameters
Throughout the two-year duration of our study, key environmental parameters were recorded at each sampling timepoint, including sea surface temperature, salinity, turbidity, photosynthetically active radiation (PAR), and oxygen saturation (Table 1). Among these, temperature, salinity, and turbidity displayed clear seasonal patterns. Sea surface temperatures ranged from 17.0–18.7°C in winter to 29.4–29.8°C in summer, with a notable seasonal amplitude of 12.4°C in 2020 and 11.1°C in 2021. Salinity peaked at 39.4–39.5 PSU during autumn, near the end of the dry season, and dropped to 38.5–39.0 PSU in winter following seasonal rainfall. Turbidity was significantly elevated in winter (FTU 2.5–2.7), likely due to increased storm activity, compared to lower values in other seasons (FTU 0.4–1.1). Day length during microplastic sampling varied seasonally as well, ranging from 10.34 hours in winter to 13.58 hours in summer. In contrast, PAR and oxygen saturation exhibited irregular, non-cyclic fluctuations, suggesting that these parameters were more strongly influenced by short-term environmental events rather than seasonal trends.
Table 1.
Seasonal environmental parameters at the time of sampling*
Diatoms on microplastics
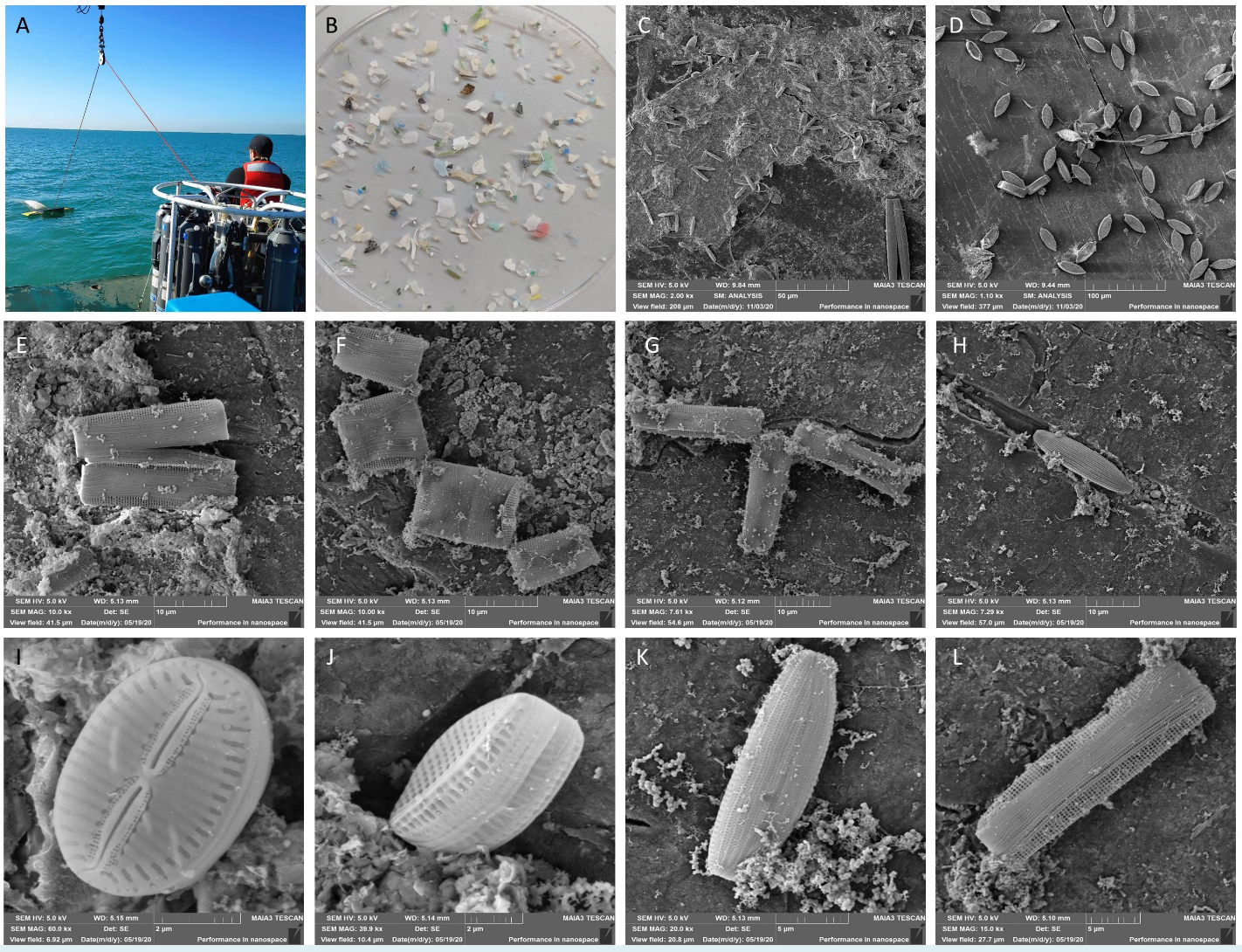
The microplastic particles (>300 µm) collected using a Manta net (Fig. 1A) exhibited a variety of colors and shapes (Fig. 1B). Scanning electron microscopy (SEM) imaging of samples collected during the winter and spring of 2020 revealed patches of bacterial biofilm with diatoms embedded within it (Fig. 1C) as well as diatoms directly attached to the plastic surfaces (Fig. 1D). Various pennate diatom species were identified (Fig. 1E–L), including individuals undergoing cell division (Fig. 1E–G).
Diatoms are relatively more abundant on microplastics compared to the surrounding seawater
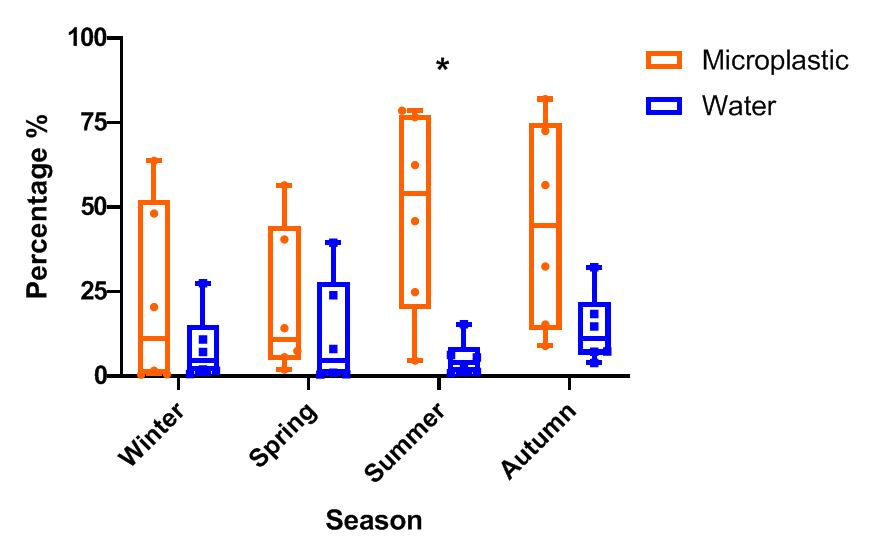
The overall taxonomic composition of eukaryotic communities, based on 18S rRNA metabarcoding, was compared between microplastic-associated samples and surrounding seawater across all seasons and samples (rarefied to 10,000 reads per sample). Diatoms were consistently more abundant on microplastics, accounting for an average of 32.15% of the 18S reads over the two-year period, compared to just 9.55% in seawater. This enrichment was observed throughout 2020 and 2021, with a statistically significant difference in summer (p < 0.05; Fig. 2). During summer, diatoms represented 48.8% of the reads in microplastic samples versus only 5.3% in seawater (averages, n = 6). A similar pattern was observed in autumn (44.6% on microplastics vs. 14% in seawater), while lower relative abundances were recorded in winter (22.5% vs. 8.3%) and spring (21% vs. 12.3%). Notably, the relative abundance of diatoms on microplastics exhibited greater variability (SD = 27.4) than in seawater (SD = 10.5), indicating higher heterogeneity among microplastic replicates (Fig. 2).
The second most abundant eukaryotic phylum on plastic surfaces after Diatomea was Retaria (20.7%), which was absent from seawater, followed by several groups uniquely enriched on plastics, including Florideophycidae (4.6%), Labyrinthulomycetes (3.1%), Cercozoa (1.8%), Ochrophyta (1.5%), and Bryozoa (1.1%). By contrast, the seawater samples were characterized by higher relative abundances of Dinoflagellata (7.6%), Protalveolata (6.5%), Prymnesiophyceae (3.9%), Cryptophyta (3.4%), and Vertebrata (2%). Fungal taxa were also more prominent in seawater, where Ascomycota (4.8%) and Basidiomycota (2.7%) exceeded their relative representation on microplastics (2.8% and 1.3%, respectively) (Davidov et al. 2024).
The microplastic contain a distinct composition of pennate diatoms
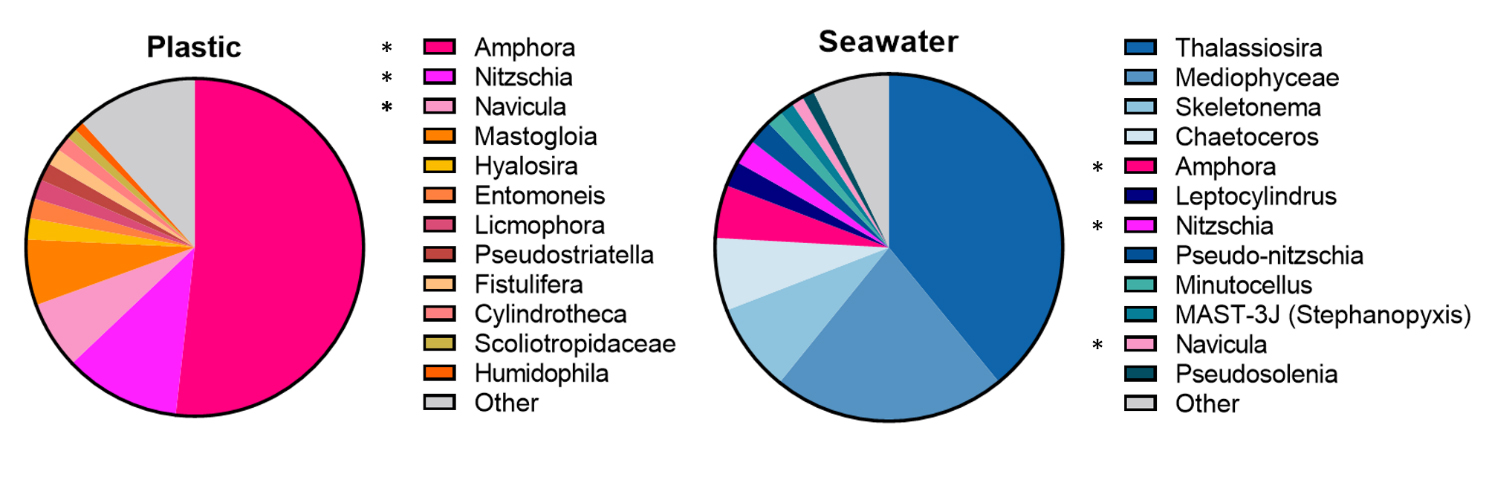
The 18S-based diatom community composition on microplastics differed substantially from that in the surrounding seawater. While raphid pennate diatoms comprised more than 98% of diatom reads on microplastics including the twelve most abundant species, seawater samples were dominated by centric diatom genera including the four most abundant species (Fig. 3). The top most abundant plastic-associated genus was Amphora (51.80% vs. 4.92% in seawater), followed by Nitzchia (11.13% vs. 2.34% in seawater). The rest of these plastic-associated benthic diatom genera had a minor representation or were absent almost completely from the seawater (e.g. Navicula). In contrary, the seawater diatom communities were dominated by Thalassiosira (93.10%), followed by Mediophyceae and Skeletonema (21.72% and 8.78% respectively) (Fig. 3).
Discussion
Floating microplastic debris serves as a novel substrate for the colonization of photoautotrophic microorganisms, particularly diatoms. The physical properties of these plastics such as low density, high durability, and, in many cases, optical transparency, make them especially favorable for diatom growth and persistence at the ocean surface. In this study we showed that diatoms thrive on microplastic in the ultra-oligotrophic waters of the Eastern Mediterranean Sea (EMS), where nutrient availability is low (Haber et al. 2022).
Our analysis showed that diatoms were the most abundant eukaryotic taxonomic group associated with floating microplastic particles. In reality, diatom representation may be even higher. As diatoms are unicellular, they will necessarily have fewer 18S rRNA gene copies than many multicellular eukaryotes, potentially leading to lower representation in metabarcoding data. This assumption is supported by our scanning electron microscopy (SEM), which showed extensive colonization of microplastic surfaces by diverse diatom taxa, predominantly raphid pennate diatoms. Diatoms were frequently observed adhering to the biofilm matrix or directly to the plastic surface, often occupying microstructural features such as pits and scratches.
The EMS marine ecosystem is characterized by extreme seasonal fluctuations in environmental parameters. In this study winter-summer temperature differences reached as high as 12.4°C. Other seasonally variable parameters were salinity with higher values in the Summer and turbidity which has increased during the Winter season. The other measured parameters (i.e. photosynthetically active radiation and oxygen saturation) did not show seasonal pattern. Results show that the diatoms were significantly more abundant on microplastic debris compared to the surrounding seawater throughout the year and particularly during the Summer season when day length, seawater temperature and solar radiation are peaking. Interestingly, the seasonal fluctuation in diatom abundance on the plastic surfaces seems to be higher than that of the planktonic counterparts. This observation may be due to the proximity of the floating microplastic to the water surface, were environmental changes are even more emphasized.
Almost all microplastic-attached diatoms that were identified in this study were raphid pennate diatoms whereas most of the planktonic diatoms belonged to centric genera. Raphid pennates possess a raphe; an elongated slit in the silica valve that enables gliding motility and is typically associated with surface-attached lifestyles. These diatoms are often found in benthic and coastal environments (Serôdio and Lavaud 2022). Their prevalence on floating microplastics suggests that such debris may serve as an important pelagic microhabitat for benthic-adapted diatom taxa, thereby expanding the known ecological niche of these organisms. Moreover, the attachment and proliferation of these traditionally benthic organisms on floating substrates may bring them into direct competition with planktonic centric diatoms.
The high abundance of diatoms on marine plastic surfaces may have significant implications for marine ecosystems. As major contributors to oceanic biogeochemical cycles, diatoms play a pivotal role in global carbon and silica cycling, both through primary production and the export of organic matter and biogenic silica to deeper ocean layers. Upon death, their heavy silicified frustules contribute to the biological pump by sinking carbon and silica to deeper ocean layers as marine snow (Tréguer et al. 2018). Diatoms’ heavy frustules may alter microplastic buoyancy from positive (floating) to negative (sinking). Dense biofilms composed of diatoms and diatom frustules have the potential to ballast microplastic particles, facilitating their downward transport into the water column. Following the senescence and decomposition of these diatom communities in the aphotic zone, the reduction in biomass may restore the positive buoyancy of the plastic particles, allowing them to resurface and be recolonized, thus forming a recurring ecological cycle.
Taken together, this two-year study highlights that floating microplastics in the EMS host distinct and diatom-rich communities, dominated by raphid pennate taxa uncommon in surrounding seawater. Seasonal patterns, especially during summer, indicate environmental conditions strongly influence diatom abundance. Together, these findings highlight novel ecological consequences of plastic colonization and underscore the need for targeted research into their long-term impacts.